Research Articles
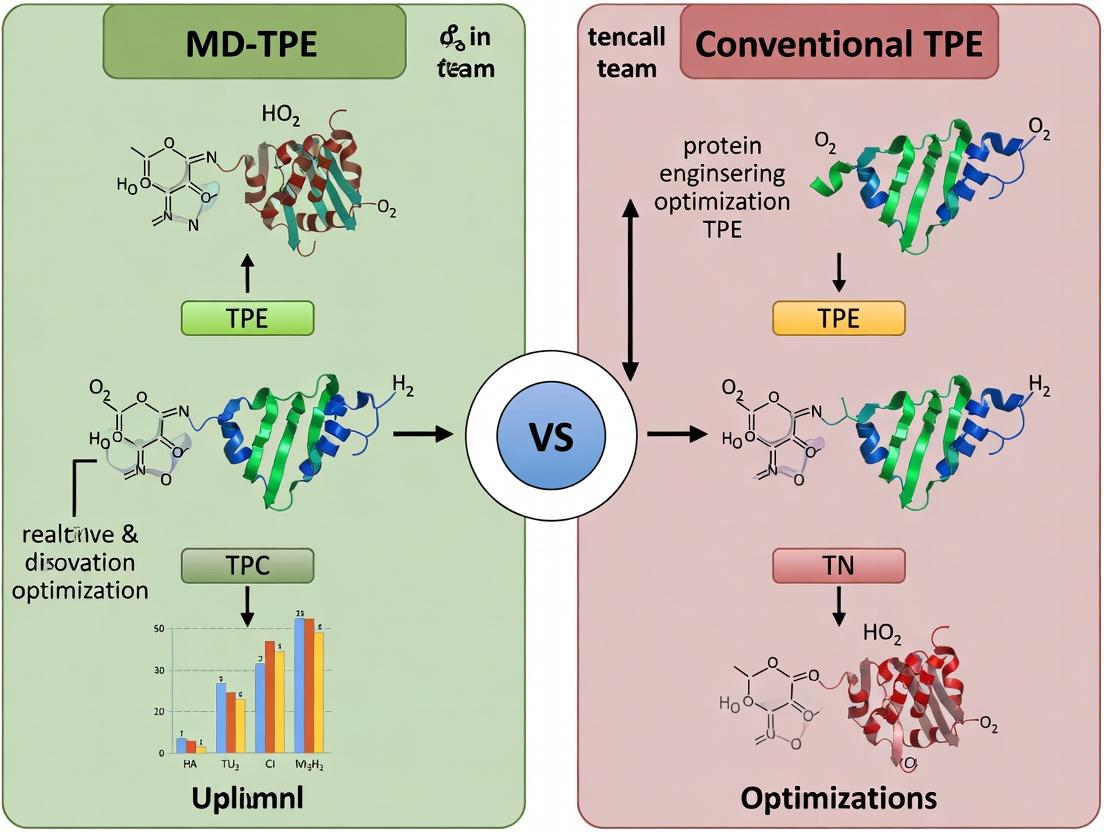
MD-TPE vs. Conventional TPE: A Comprehensive Guide to Safer, Smarter Bayesian Optimization in Drug Discovery
This article provides a targeted analysis for drug development researchers on the application of Multivariate Deep Tree-Structured Parzen Estimator (MD-TPE) versus conventional Tree-structured Parzen Estimator (TPE) for hyperparameter optimization.
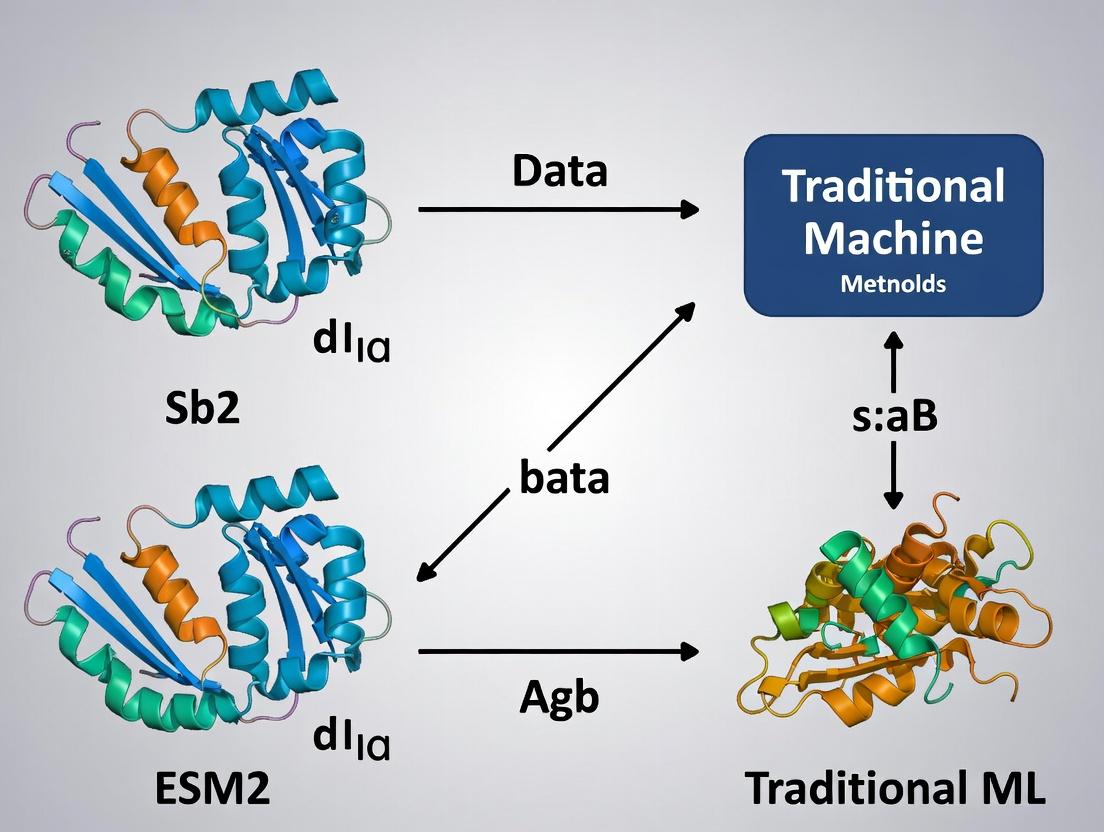
ESM-2 vs. Traditional ML: The AI Revolution in Protein Function Prediction for Biomedical Research
This article provides a comprehensive comparative analysis of ESM-2 (Evolutionary Scale Modeling-2) transformer models and traditional machine learning methods for protein function prediction.
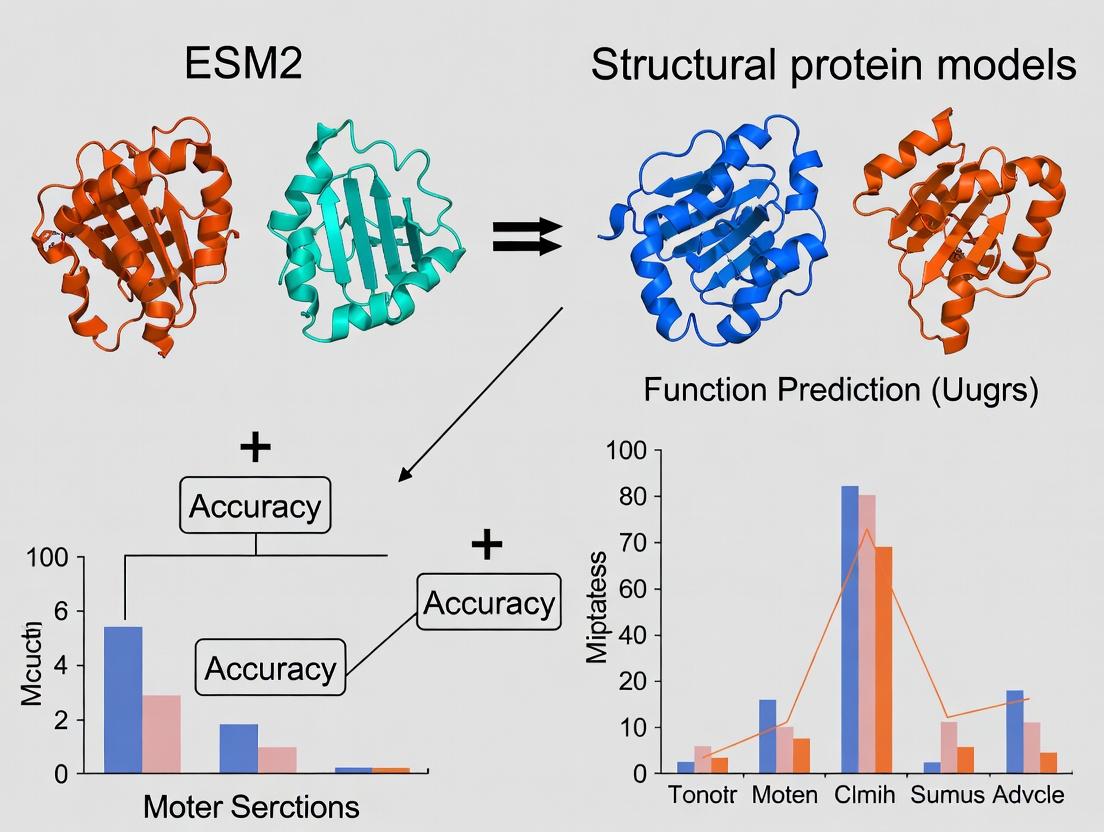
ESM2 vs. AlphaFold2: Benchmarking Protein Function Prediction Accuracy for Drug Discovery
This article provides a comprehensive comparison of the function prediction capabilities of the revolutionary language model ESM-2 and leading structural models like AlphaFold2.
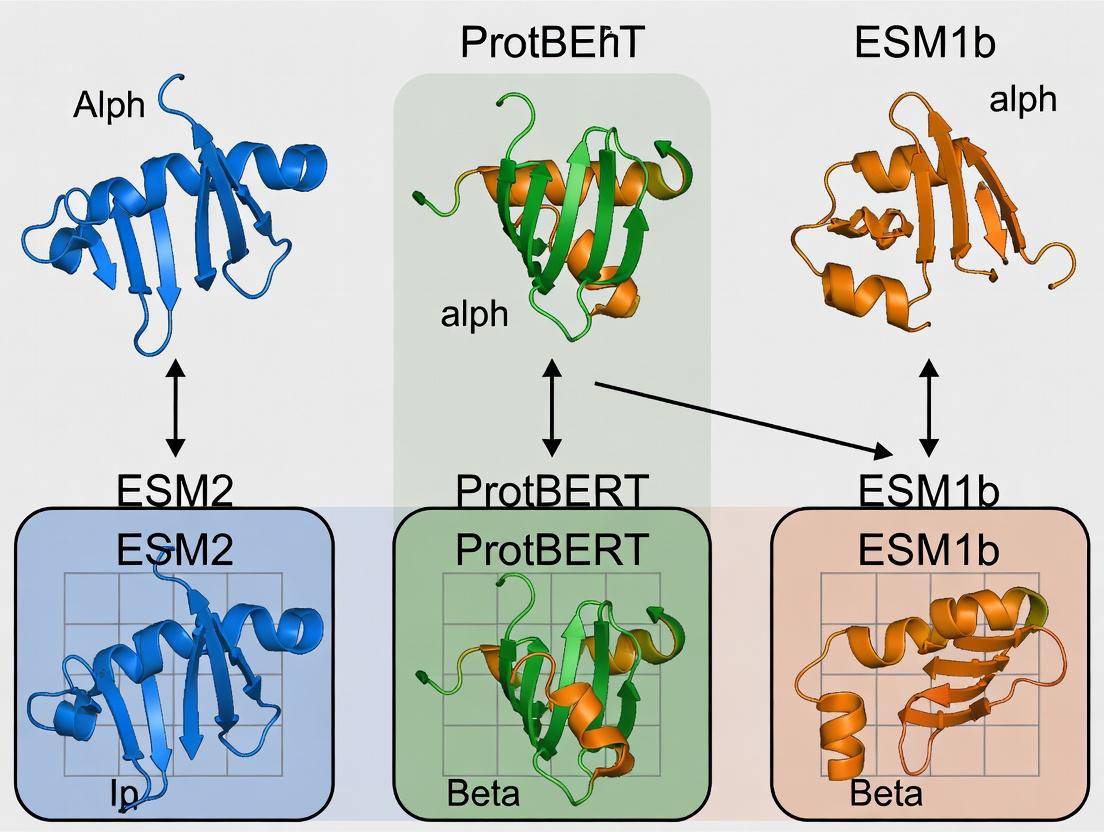
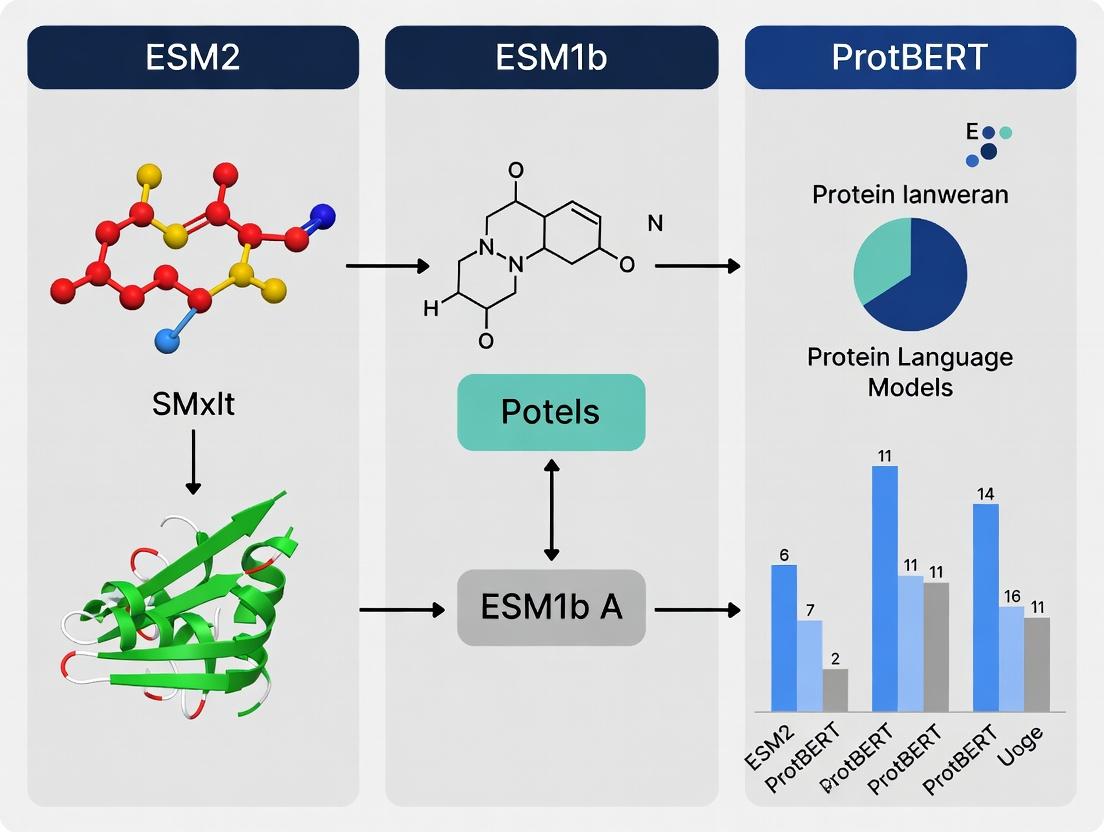
EC Number Prediction Showdown: A Deep Dive into ESM2 vs. ProtBERT vs. ESM1b Performance
This comprehensive analysis compares three leading protein language models—ESM2, ProtBERT, and ESM1b—for the critical task of Enzyme Commission (EC) number prediction.
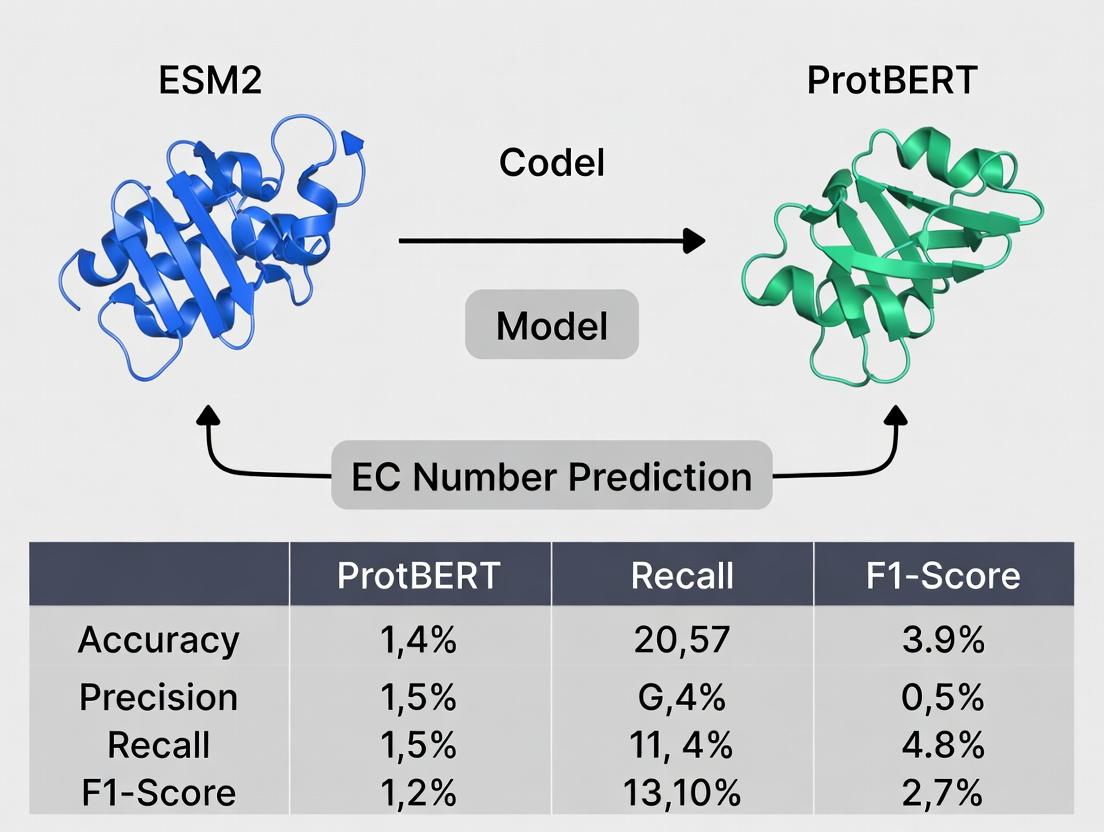
ESM2 vs ProtBERT: A Comprehensive Performance Analysis for EC Number Prediction in Protein Function Annotation
This article provides a detailed comparative analysis of two state-of-the-art protein language models, ESM2 and ProtBERT, for the critical task of Enzyme Commission (EC) number prediction.
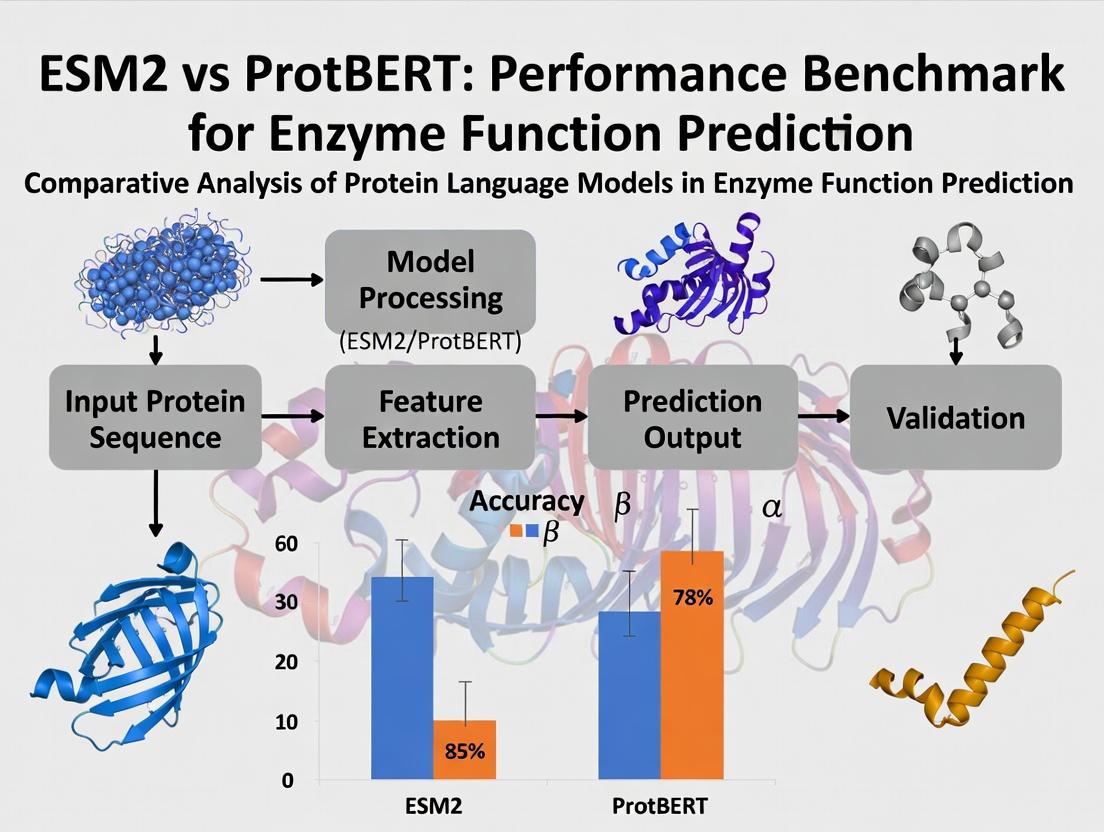
ESM-2 vs ProtBERT: Benchmarking Deep Learning Models for Accurate Enzyme Function Prediction
This article provides a comprehensive performance benchmark for two state-of-the-art protein language models, ESM-2 and ProtBERT, in predicting enzyme function.
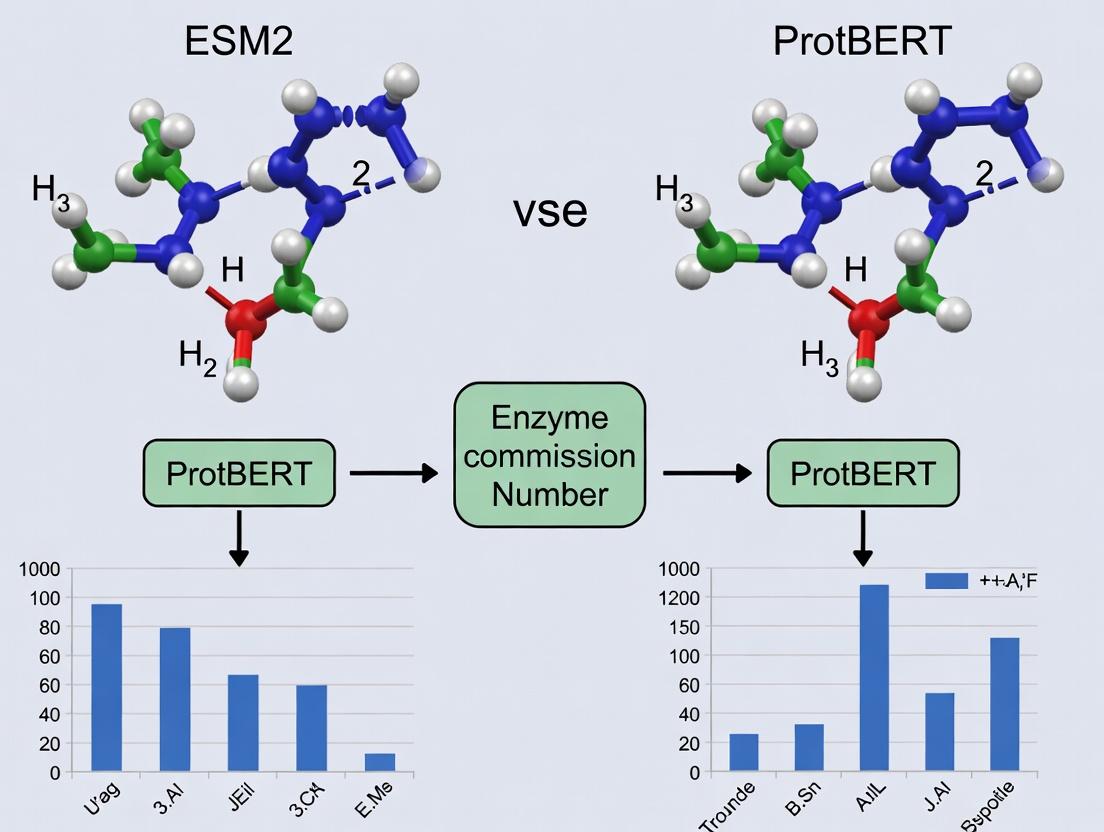
ESM2 vs. ProtBERT: Which AI Model is Superior for Enzyme Function Prediction?
This article provides a comprehensive comparison of two state-of-the-art protein language models, ESM-2 and ProtBERT, for the critical task of predicting Enzyme Commission (EC) numbers.
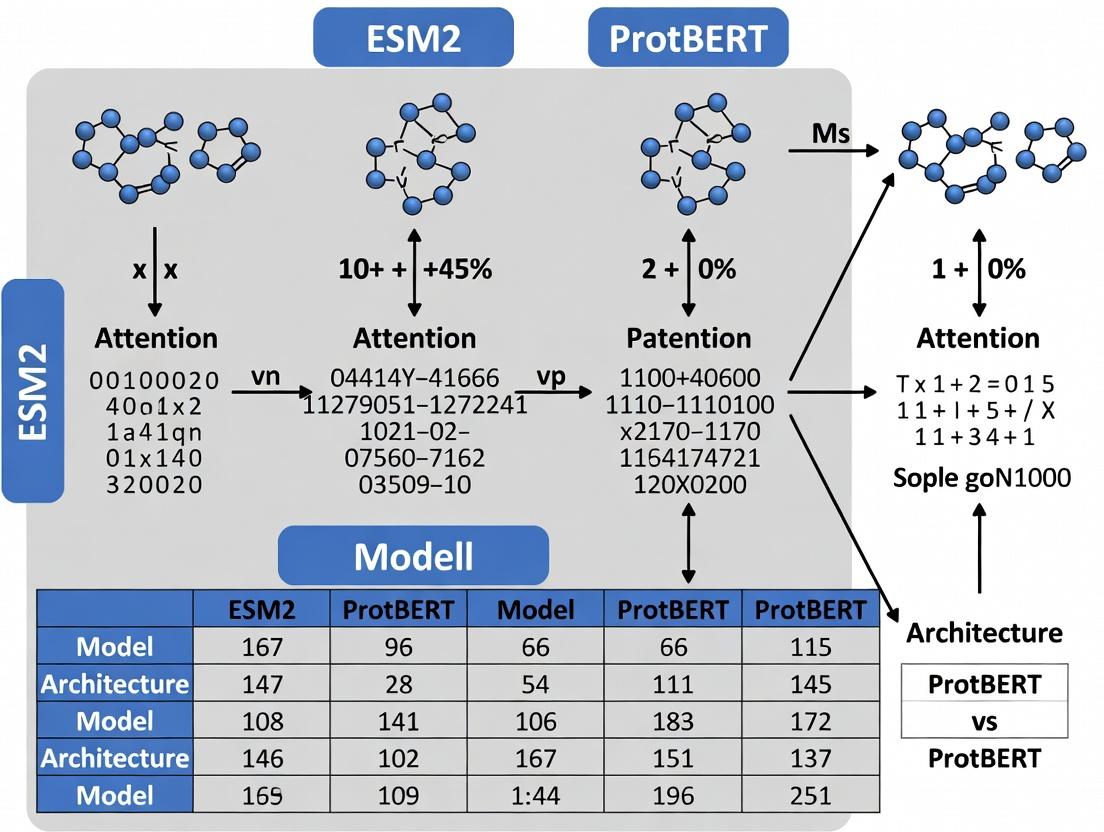
ESM2 vs ProtBERT: A Comprehensive Guide to Architecture Differences and Protein Language Model Applications
This article provides a detailed comparative analysis of two leading protein language models, ESM-2 and ProtBERT.
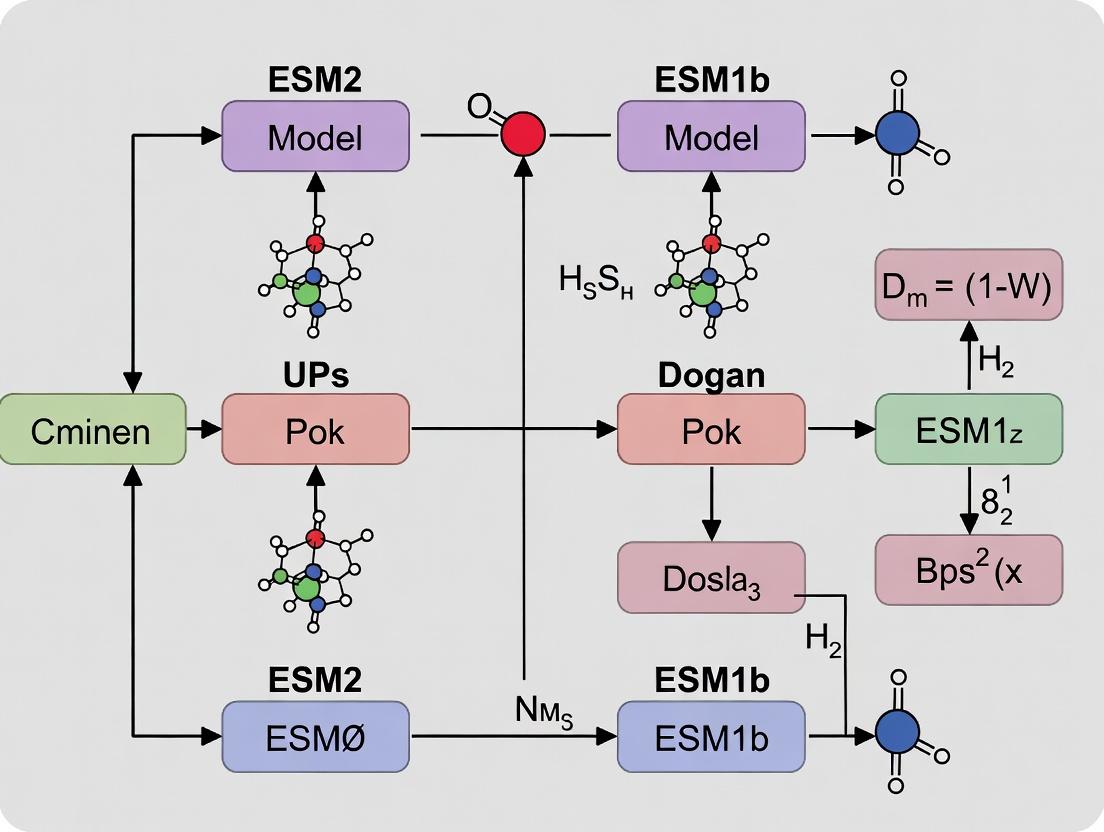
ESM2 vs ESM1b vs ProtBERT: A Comparative Guide for Biomedical AI Researchers
This article provides a comprehensive analysis and practical comparison of three leading protein language models—ESM2, ESM1b, and ProtBERT.
ESM2 vs ESM1b: A Comprehensive Performance Comparison for Biological Tasks in Drug Discovery
This article provides a detailed comparative analysis of the ESM2 and ESM1b protein language models, focusing on their performance, applications, and practical utility in biological research and drug development.